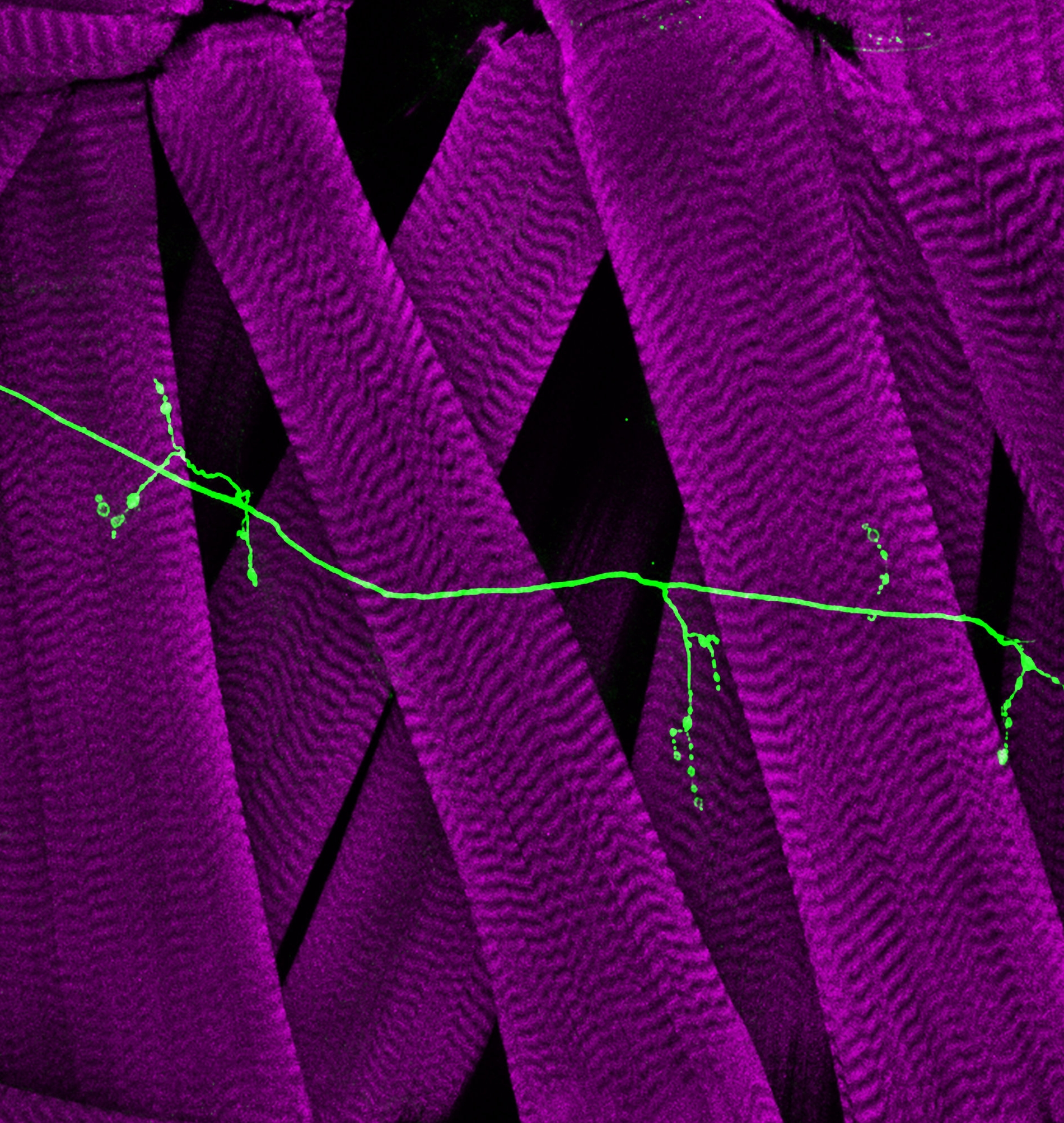
How is neural circuit specificity achieved?

Overview
Neurons communicate with each other at specialized junctions called synapses. Understanding when, where, and how these synapses develop will shed light on everyday behaviors including memory acquisition and retrieval and motor movements.
To properly transmit information, synaptic connections must be highly specific and only form between appropriate neuronal partners. These “special connectional relationships” were described in the late 1800’s by Santiago Ramòn y Cajal as “protoplasmic kisses that appear to constitute the final ecstasy of an epic love story.” As romanticists, members of the Carrillo lab are interested in understanding this love story and how it blossoms over time.
1) Finding the right partner(s): Synaptic connections are highly stereotyped, facilitating our analysis of genes that instruct the assembly of circuits. We use the Drosophila nervous system to uncover “lock-and-key” mechanisms mediated by cell surface proteins.
2) Growing together: After finding the right partner(s), synapses expand to maintain proper synaptic activity. We explore how many of the same cell surface proteins that establish the synaptic connections subsequently drive the expansion of the same synapses.
3) Compensation upon loss of a neighbor: Synaptic plasticity allows neurons to strengthen or weaken their activity in order to compensate for changes in function. We are interested in understanding how one synaptic partner detects and responds to perturbations in a neighboring neuron.
Overall, we utilize genetic, biochemical, and functional approaches to uncover fundamental programs that mediate nervous system development.
Updates
March 2026: Happy spring equinox! Nowruz mubarak! (happy Persian new year), happy Gudi Padwa (Marathi new year) , Eid Mubarak! (happy end of Ramadan)
January 2026: Congratulations to Camille Casiño on joining the lab as our new research analyst! We're so excited to have you aboard the Carrillo Cruise.
January 2026: Welcome to Austin Li who will be rotating in the lab this quarter! We are so happy to have you with us!
'Tis the season! From the Carrillo lab: ¡Feliz Navidad! Maligayang Pasko! Happy Kwanzaa! Shab-e yalda mobarak! Happy Holidays!!
Its grant applicaiton season! Good luck to Robert, James, Idris, and Luciano on their respective grant submissions.
Happy Thanksgiving! We hope you enjoyed getting together with loved ones and sharing your gratitudes!
November 2025: Renee is on her way to present a poster at the Society for Neuroscience conference in San Diego! You're gonna kill it!
October 2025: Feliz Dia de los Muertos / Happy Halloween! Spooky season is a favorite around the Carrillo Lab.
October 2025: Happy indigenous peoples day!
October 2025: Huge congratulations Ellen on being awarded "best poster" for a postdoc/senior scientist at the Midwest Drosophila Conference! We're so proud of you!!!
October 2025: Ellen is on her way to CSHL to attend the Neurobiology of Drosophila conference! Good luck and have fun!
October 2025: Renee is on her way to give a poster at the Janelia Biology and Engineering of the Cell Surface Conference in Ashburn, VA!! Safe travels and good luck!
October 2024: Congratulations to Hunter Richardson for joining the lab! We're thrillled to have you, welcome to la familia!
September 2025: The Carrillo Lab is on its way to its first annnual Lab Retreat! We're looking forward to coming together as a team and brainstorming for the future!
August 2025: Ellen is on her way to TA the KITP Neurophysics of Sensing in Motion, Systems Neuroscience Summer Course! We'll miss you!
August 2025: It is Matilde's last day! We've loved having you in the lab, we wish you well in your future endeavors!
July 2025: It is Omr Kassem's last day in the lab. He will be beginning his graduate adventures in Germany! We are sad to see you go but happy to watch you thrive! All the best luck, Omr!!
June 2025: Robert, James Ashley, Vasudha Aher, and Luciano Simonetta are at Cold Spring Harbor Laboratory for the course, Drosophila Neurobiology: Genes, Circuits & Behavior. It’s Robert’s final year as course co-director and both Vasudha's and Luciano's first time as TAs. Make the most of it, y'all!
June 2025: Matilde Grosso a DARN (Diversity Access to Research in Neuroscience) fellow from Universidad de Puerto Rico Rio Piedras has joined the team for the summer. Excited to have you!
May 2025: The Carrillo lab is running a two-day workshop for high school students at Back of the Yards College Preparatory High School. Big shout out to Rio Salazar and Luciano Simonetta for leading it, and thank you to everyone who came out to help.
March 2025: Huge shout out to Rio Salazar, Hunter Richardson and Idris Ayantoye for running the Shamrock Shuffle 8K!! You guys are powerhouses!
October 2025: Vasudha is heading to the MBL for the Embryology course! Have so much fun!
July 2024: Cogratulations to Ellen Lesser for joining the lab as our first official post-doc! We're so excited to have you and your expertise. Welcome to the family!
July 2024: On our way to Lucca, Italy for the Gordon Research Conference in Neurodevelopment! Rio Salazar is giving a research talk at the Gordon Research Symposium for graduate students and Renee Zhang is presenting a poster at the GRC. Buon viaggio, ragazze!
July 2024: NIH Multi-PI R01 with Engin Özkan begins. Our great collab continues!
July 2024: Robert, James Ashley and Parisa Tajalli are at Cold Spring Harbor Laboratory for the course, Drosophila Neurobiology: Genes, Circuits & Behavior. It’s Robert’s second year as course co-director and Parisa’s first time as a TA. James ran a stellar e-phys demo!
June 2024: Congratulations to Vasudha Aher for passing her prelim unconditionally! You’re a baller!
June 2024: Luciano Simonetta receives an NIH diversity supplement. You’re killin’ it!
June 2024: Natalia Jiménez a DARN (Diversity Access to Research in Neuroscience) fellow from Universidad de Puerto Rico Rio Piedras has joined the team for the summer. Excited to have you!
June 2024: Welcome to the new 2024 NEETO fellows and their teacher Paul Khairallah from Back of the Yards College Prep! Luciano Simonetta is rocking it in his second round as the graduate student mentor. The Fellows arrive on Luciano’s birthday. ¡Feliz cumple, Lu!
June 2024: Welcome Hunter Richardson, MSTP student, rotating in the lab!
May 2024: Congratulations to Vasudha Aher for joining the lab! We’re excited to have you, welcome to la familia.
May 2024: As a follow-up to last summer’s NEETO program the Carrillo lab is running a two-day workshop for high school students at Solorio High School. Big shout out to Luciano Simonetta for leading it, and thank you to everyone who came out to help.
April 2024: Robert takes on a new DEIJ leadership role as Associate Dean for DEI – Basic Sciences.
March 2024: Robert is giving a seminar at The University of New Mexico. We miss you!
March 2024: It’s spring equinox and Persian new year, norooz mobarak!
March 2024: Robin Mitchell joins the lab! So excited to have you on the team.
Feb 2024: Happy Lunar New Year! Year of the dragon!
February 2024: Robert is at USC giving a talk in the Department of Neurobiology. Hope he’s re-upped on vitamin D!!!
NEETO visits Kennedy High School on the Southwest Side of Chicago. Kudos to Parisa Tajalli for leading the two-day workshops. Big thanks to everyone who participated! We gorged on Birria after on the way back to UChicago!
February 2024: Robert gets tenure! So excited we don’t have to move to sunny California!
January 2024: Committee on Neurobiology student Robin Mitchell rotates in the lab. Welcome, Robin!
December 2023: Christmas and winter solstice are around the corner: Feliz Navidad, Frohe Weihnachten, Shab-e yalda mobarak, and Happy Holidays from the Carrillo lab!
December 2024: Lots of gluhwein and hot cocoa at the christkindlmarket … we didn’t freeze to death.
November 2023: Happy indigenous peoples day! We had fun celebrating with the lab… James robbed us in poker while we enjoyed all the great pot-luck food.
September 2023: Carrillo Lab hosts keynote speaker Marc Freeman from OHSU Vollum Institute at the UChicago Neuro retreat. Renee and Rio have stellar posters!
September 2023: Congratulations to Luciano Simonetta on 're'joining the lab! Glad that your rotations brought you back to la familia!
September 2023: Renee is awarded the NIH F31 Fellowship! Congratulations!!!!
Robert is off to soak up the sun in Puerto Rico for the Ivey Plus recruitment
Taliana Salcedo Rosado, DARN fellow from Universidad de Puerto Rico Bayamón presents at the Leadership Alliance end of Summer Symposium. We’ll miss you!
August 2023: Chris presents at the 2023 NESSTP Symposium
August 2023: Dr. Meike Lobb-Rabe gives a stellar defense. We'll miss you Meike! Can't wait to see what amazing things you do next!
July 2023: Taliana presents at the 2023 Leadership Alliance National Symposium.
July 2023: Rio and Robert are awarded the HHMI Guilliam Fellowship. Congratulations!!!!
July 2023: Robert is heading-up the CSHL Drosophila Neurobiology Course this year. James is leading a section on NMJ development and Rio showing off her killer embryo dissections as our favorite TA!
June 2023: Taliana Rosado Salsedo joins us from the Universidad de Puerto Rico en Bayamon as a DARN fellow participating in Leadership Alliance. Welcome, Taliana!
June 2023: This year's Neuroscience Early Traning Opportunity (NEETO) fellows and their teacher, Gina Vittoria, arrive from Kenedy High School. It's going to be a jam-packed 2 two weeks!
June 2023: Rio and Parisa present a poster at the UChicao DEIJ retreat. Reach out if you want to get involved in our programs: NEETO, DARN, UNDO, DNUFO
'Tis the season! From the Carrillo lab: ¡Feliz Navidad! Frohe Weihnachten! Happy Kwanzaa! Shab-e yalda mobarak! Happy Holidays!
October 2022: Rio Salazar presents a poster and Renee Zhang gives a talk at the MGCB retreat. Great job!
October 2022: The Carrillo lab is off to the MGCB retreat. First in-person retreat since the pandemic started!
October 2022: Rio and Robert are off to Puerto Rico to recruit graduate students and summer research programs. Buen viaje, birng us back some coquito!
October 2022: Meike Lobb-Rabe and Katie DeLong’s paper is out! Great job team!
Dpr10 and Nocte are required for Drosophila motor axon pathfinding
October 2022: Aurora Ferrell and Chris Cardenas join the lab!
September 2022: CMB student Karen Velez rotates in the lab. Welcome, Karen!
September 2022: Meike Lobb-Rabe gives a poster at the Molecular Mechanisms of Neural Connectivity.
September 2022: Congrats to Parisa Tajalli on being awarded a spot on the Developmental Biology Training Grant!
August 2022: Meike Lobb-Rabe gives a talk at the Neural Development Gordon Conference
August 2022: Congratulations to Rio Salazar for passing her qual!
August 2022: MSTP student Idris Ayantoye rotates in the lab. Welcome, Idris!
July 2022: Robert’s DNUFO students present their work at the summer symposium!
June 2022 Luciano Simonetta presents at the PREP summer symposium
June 2022: DRSB student Parisa Tajalli joins the lab. Welcome to the familia, Parisa!
June 2022: Darion Estevez joins the lab. Welcome to the familia, Darion!
June 2022: Yupu Wang graduated. We’ll miss you, Yupu. Can’t wait to see what amazing things you do next at Janelia!
May 2022: Dr. Yupu Wang brought a tear to Robert’s eye as he gave a stellar defense. Congratulations, Yupu!
April 2022: PREP student, Luciano Simonetta accepts offer to join the DRSB graduate program at UChicago!
January 2022: DRSB student, Parisa Tajalli Tehrani Valverde rotates in the lab!
November 2021: The Carrillo Lab was featured by the UChicago Faculty Development Program!
A neurological love connection | UChicago Faculty Development Program | The University of Chicago
August 2021: Welcome Luciano Simonetta! Luciano is a PREP Scholar and will be with us for the upcoming academic year.
August 2021: Congrats to Renee Zhang on being awarded a spot on the Molecular and Cellular Biology Training Grant!
July 2021: Welcome to our new research technician Stephanie Dunning!
July 2021: We have two new(ish) lab members! Renee Zhang (DRSB) and Rio Salazar (CMB) have made it official and are now grad students in the Carrillo lab! The group is growing 😊
June 2021: Congrats to Renee and Rio on passing their qualifying exams unconditionally! Ohhh yeah!
June 2021: Super happy to announce that the Carrillo lab was awarded an NIH R01!! Thanks to all the lab members for the hard work, Ilaria Rebay for the long hours helping to edit, and the reviewers and NIH for their continued support!
March 2021: Renee Zhang and Rio Salazar join the lab as Spring rotation students! Glad to have you both onboard!
February 2021: Excited to announce that the Carrillo lab was awarded an NSF CAREER award!!! Congrats to all lab members and everyone who helped along the way. Special shout out to Chip for the extensive help and Jean! Also, thank you to the grant reviewers and NSF for their support!
January 2021: Yupu got the best poster award from MGCB retreat! Congrats!
January 2021: Our paper was selected for the Journal of Neuroscience cover! Major shout out to Purujit Chatterjee, our former undergrad, for the beautiful artwork!
December 2020: Yupu’s papers is in Journal of Neuroscience! Congrats to Yupu and collaborators (Meike, James, Veera, and all who helped)!
October 2020: Meike is awarded an F31 Fellowship! CONGRATS!!
September 2020: CMB student, Annie Green, rotates in the lab!
September 2020: Undergrad student, Iris Berman, joins the lab!
June 2020: Our manuscript about synaptic plasticity paper is available on BioRxiv!
May 2020: Collaboration with the Hecksher lab available on Elife! (see Publications)
October 2019: Yupu is awarded a Second-year International Student Fellowship from BSD!! Ohhh yeah!
September 2019: Congrats to Luke Shackleford (REU student) for being selected to present at the national REU symposium in DC!
September 2019: Meike wins best poster presentation at UChicago Neuro retreat!
June 2019: REU student, Luke Shackleford, joins the lab for the summer!
June 2019: DNUFO students, Katherine Hou and Nico Cordero, join the lab!
June 2019: Our second grad student, Yupu Wang, joins the lab!
April 2019: Collaboration with the Vanderzalm lab available in Developmental Neurobiology! (see Publications)
February 2019: The lab's first paper available on eLife! (see Publications)
January 2019: Collaboration with the Özkan lab available on eLife! (see Publications)
January 2019: CMB student Sarah Yde rotates in the lab!
September 2018: DSRB student Yupu Wang rotates in the lab!
July 2018: CMB student Noah Peña rotates in the lab!
June 2018: Our first grad student, Meike Lobb-Rabe, joins the lab!
June 2018: Undergrads Katie DeLong and Gabriel Rojas-Bowe join the lab for the summer!
March 2018: Grad Students Meike Lobb-Rabe (CMB) and Viola Nawrocka (BMB) rotate in the lab!
January 2018: Grad Students Craig DeValk (BMB) and Erica Mezias (CON) rotate in the lab!
October 2017: Staff Scientist James Ashley joins the lab!
September 2017: CON student Marie Greaney rotates in the lab!
July 2017: Lab Tech Violet Sorrentino and Undergrad Veera Anand join the lab!
June 2017: Our first undergrad, Purujit Chatterjee, joins the lab!
May 2017: The very first lab member, Chloe Weisberg (Lab Tech), joins the lab!
March 2017: The lab is awarded a BSD Diversity & Inclusion Faculty Grant!
January 16, 2017: The Carrillo Lab is open for business! (not fake news!)
Follow us on X @NufoTheFlyGuy